|
Cytoscape offers out-of-the-box integration with public network databases such as NDEx.The plugin enables the user to use the available file formats for importing and exporting graph workspaces from SAP HANA. GraphML, SIF, GML and many more) and also has import wizards for general purpose file formats such as CSV or Excel. Cytoscape integrates with a variety of network data formats (e.g.With a few clicks, you can visually explore graph data stored in SAP HANA. Cytoscape provides an easy-to-use rich client to visualise, explore and understand network/graph data.Even with only basic connectivity, users will immediately gain advantages when working with the plugin: 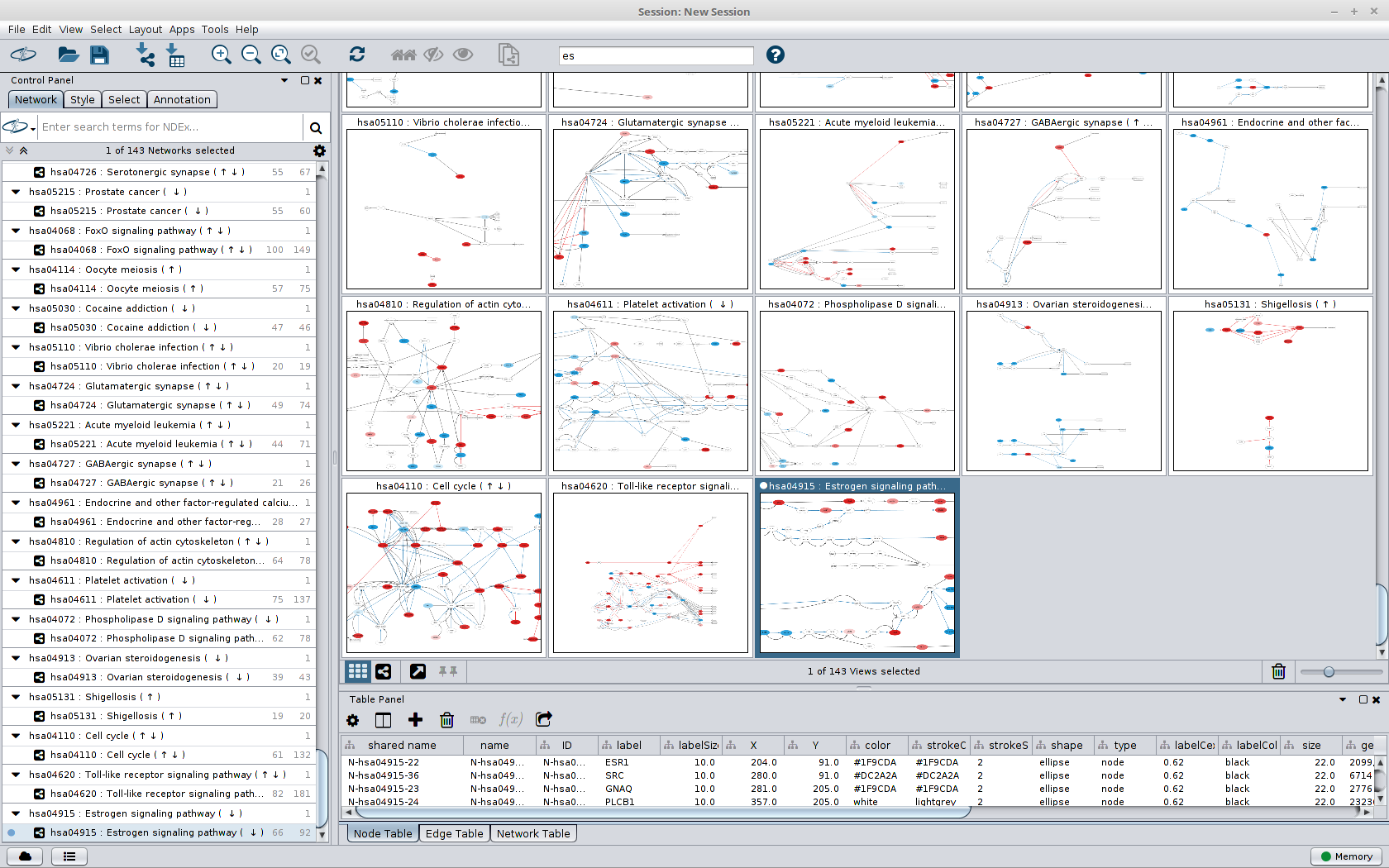
The plugin shall provide basic connectivity between Cytoscape and Graph Workspaces in SAP HANA. (Time feels infinite when 4 kids leave the house – so this ended up being a few thousand lines of code) cyrest_get ( 'networks.Recently, my family was out for a couple of days, which gave me time to kick start a little spare-time project: An SAP HANA plugin for the open source network analysis client Cytoscape. Returns: dict: ] """ cy_networks_suids = commands. Default is and the latest version of the CyREST API supported by this version of py4cytoscape. Args: network (SUID or str or None): Network name or SUID of the network that you want set as current base_url (str): Ignore unless you need to specify a custom domain, port or version to connect to the CyREST API. def set_current_network ( network = None, base_url = DEFAULT_BASE_URL ): """Selects the given network as "current". py4cytoscape_sandbox import get_abs_sandbox_path def _init_ ( self ): pass py4cytoscape_tuning import MODEL_PROPAGATION_SECS, CATCHUP_NETWORK_SECS, CATCHUP_NETWORK_TIMEOUT_SECS from. import sandbox # Internal module convenience imports from. General network functions # - # External library imports import sys import time import warnings import pandas as pd import igraph as ig import networkx as nx # Internal module imports from. 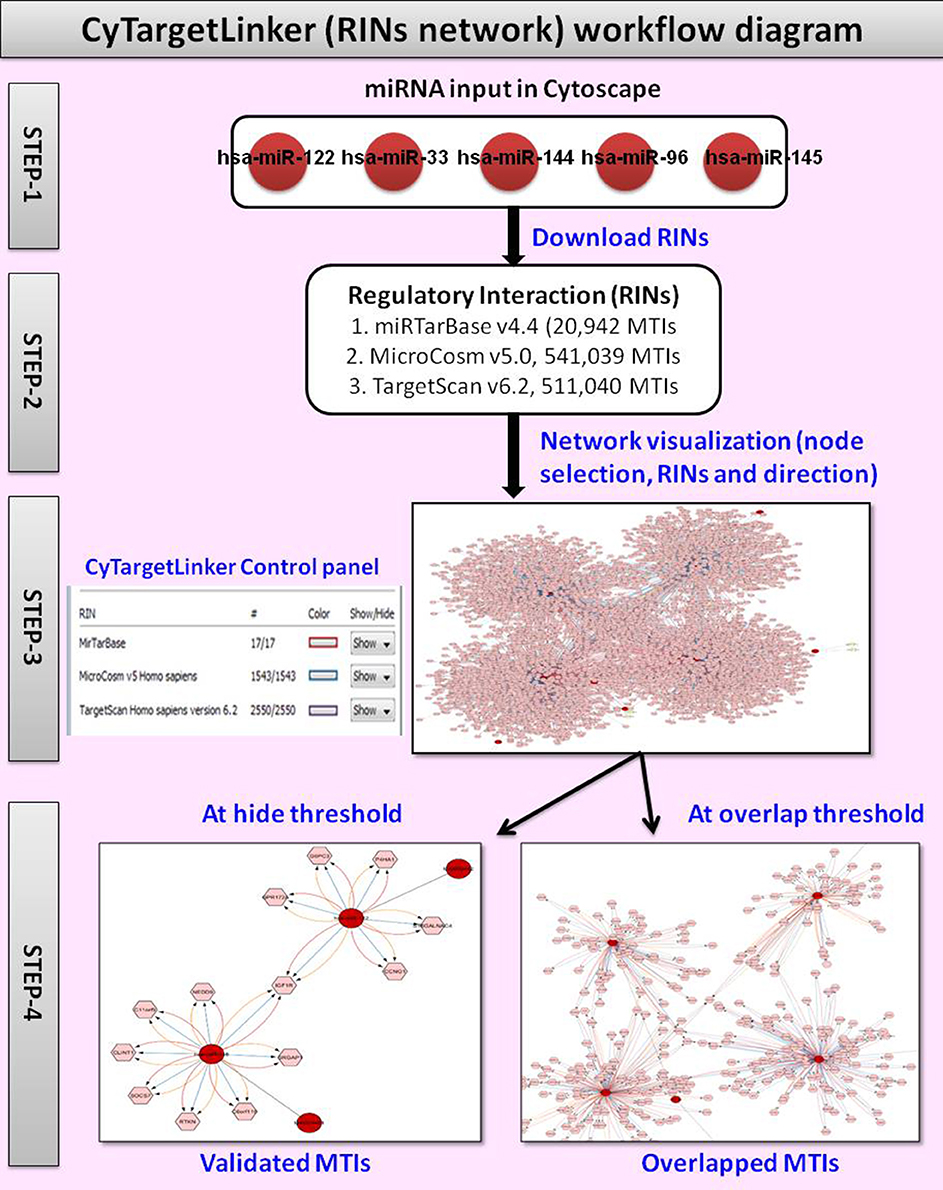
IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE. THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. 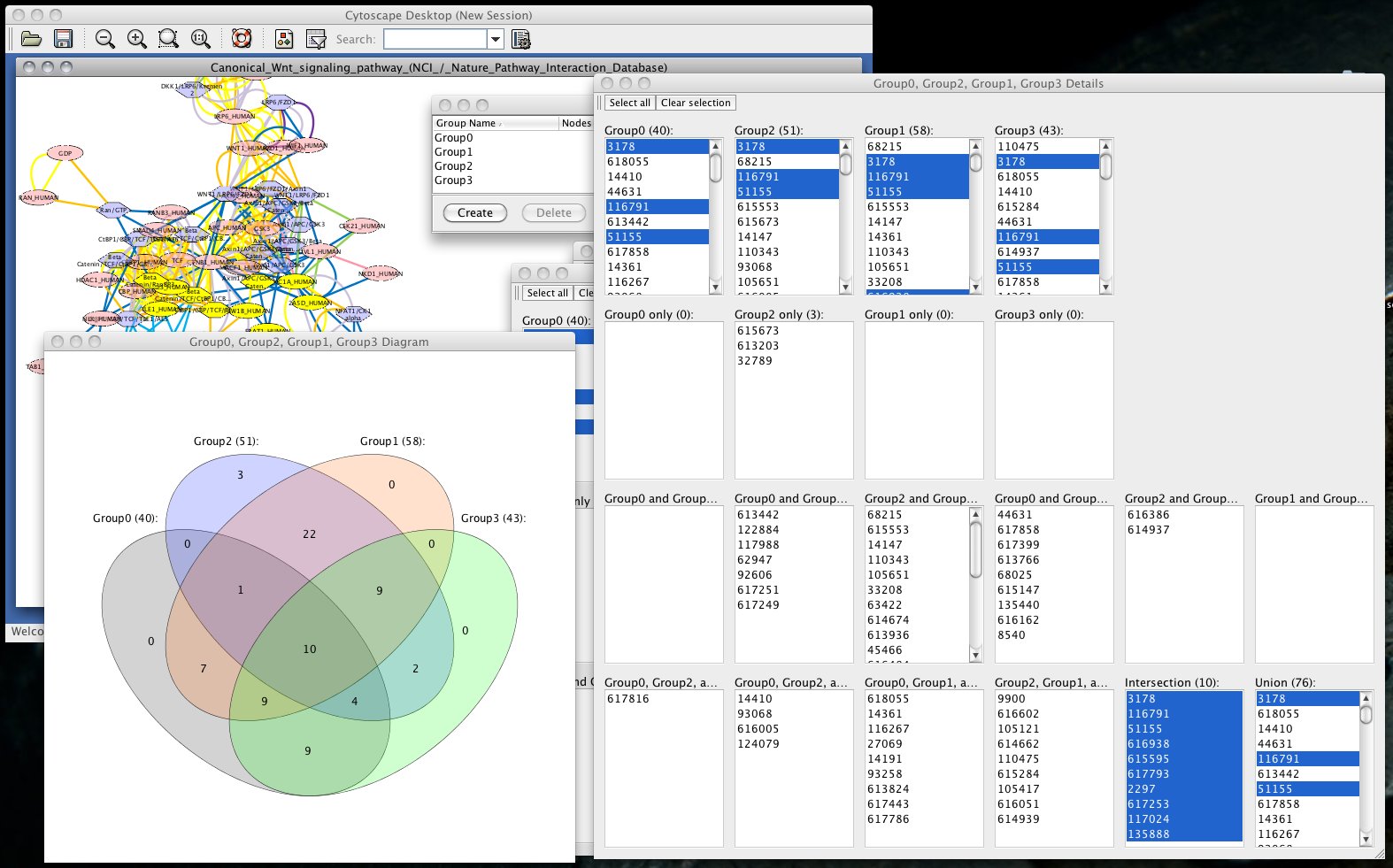
""" """Copyright 2020-2022 The Cytoscape Consortium Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions: The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software. See the ``Utils`` section for functions that convert node and edge names to SUIDs, and vice versa. Internal functions Note: See the ``Network Selection`` section for all selection-related functions. Includes all functions that result in the creation of a new network in Cytoscape, in addition to funcitons that extract network models into other useful objects. # -*- coding: utf-8 -*- """Functions for NETWORK management and retrieving information on networks, nodes and edges.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
